Production of oligonucleotide second messengers to activate antiviral immunity is an ancient and evolutionarily conserved defense mechanism. In eukaryotes this includes cGAS-STING, and in prokaryotes this comprises related enzymes in the cyclic oligonucleotide-based antiphage signaling system (CBASS) and Type III CRISPR systems.1 In all known systems, antiviral cyclases are tightly regulated, becoming active upon the detection of viral infection. Here, the authors characterize the Panoptes prokaryotic immune system, which comprises a cyclase (minimal CRISPR polymerase, mCpol) and antiviral effector (2TMβ).2,3 While other antiviral cyclases remain inactive until infection, the structure of mCpol reveals a striking deviation: it produces cyclic oligonucleotides without external stimulus. This structure, in concert with biochemical characterization, reveals that Panoptes acts through a novel signaling paradigm where the oligonucleotide second messenger is constitutively produced, repressing the immune response. This signaling logic allows for Panoptes activation in response to viral encoded anti-defense proteins that aim to subvert immunity by depleting antiviral second messengers.
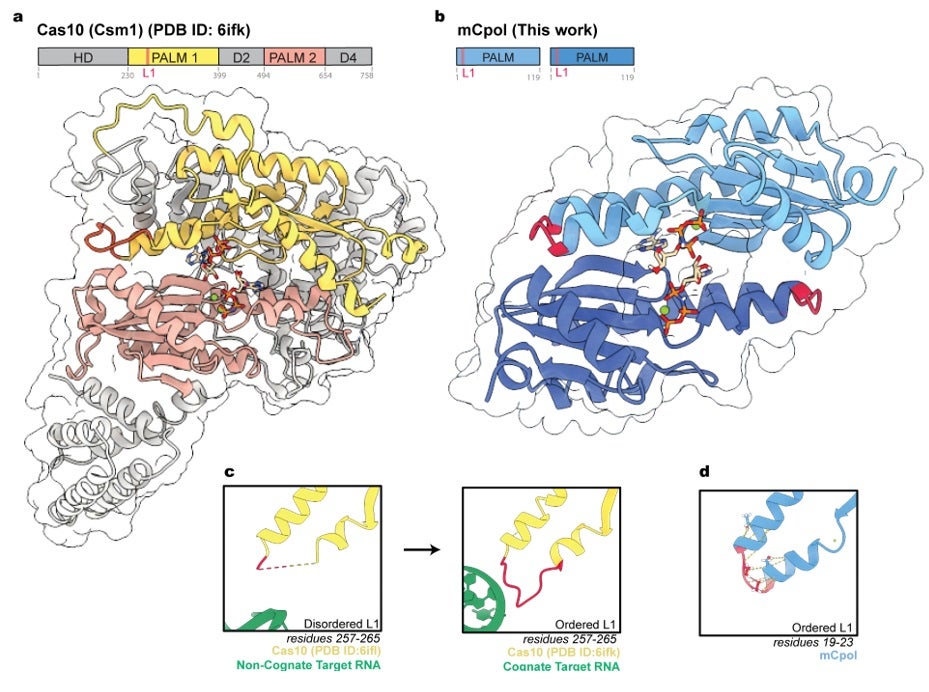
A previous comparative genomics analysis identified this putative cyclase, mCpol, as a relative of the characteristic enzyme of Type III CRISPR systems, Cas10.4 Cas10 is a cyclase that produces oligonucleotide messengers upon binding of a target RNA, indicating the presence of virus.5,6 Allosteric regulation of Cas10 activity provides control over cyclic oligonucleotide synthesis, fine-tuning it in response to viral infection.7 The Loop L1 region of Cas10 (residues 19-23 of SthCsm1) is disordered prior to target RNA binding. However, in the presence of target RNA this loop becomes ordered, increasing cyclic oligonucleotide synthesis to activate an antiviral response.7
This work examined a putative antiviral system, Panoptes, containing mCpol and an antiviral effector. Incubation of recombinant mCpol with ATP showed the formation of cyclic oligonucleotide products, primarily 2’3’-c-diAMP, indicating that mCpol was an active cyclase. This activity paralleled that seen by Cas10 in previous studies.5,6 However, it did not seem to require additional stimulus such as target RNA binding. To gain insight into this divergent mechanism of regulation, a high-resolution crystal structure of mCpol bound to an ATP analog (ApNHpp) was determined at 2.3 Å resolution using data collected on SSRL BL12-2. The structure of the Panoptes cyclase, mCpol, and corresponding biochemical analysis show that this enzyme folds into a single Palm domain which dimerizes to form a pocket where two ATP analogs are bound. This dimer structure is structurally homologous to the two Palm domains that comprise Cas10,8,9 allowing for the comparison of regulatory features between the two cyclases.
Examining the region of mCpol homologous to Loop L1 in Cas10 revealed a divergence in this allosteric regulatory region. The disordered loop in Cas10 was replaced by a highly ordered turn in mCpol, anchoring this catalytic-adjacent region. Additionally, mCpol lacked structural homology to other regulatory regions of Cas10.8 This insight was consistent with the lack of RNA-dependence and constitutive in vitro activity of mCpol. However, it was unclear how constitutive oligonucleotide synthesis could lead to a response specific to viral infection.
Due to mCpol’s constitutive activity, it was then hypothesized that the system containing mCpol operated through oligonucleotide mediated negative regulation, where the messenger suppressed the effector protein. Cell-based and biochemical assays demonstrate that the presence of 2’3’-c-diAMP produced by mCpol suppresses the active, oligomeric form of the associated effector protein thereby suppressing Panoptes activity. This inversion of known oligonucleotide signaling logic allows the system to become active upon oligonucleotide depletion. Notably, the bacterial viruses sensitive to Panoptes all encode anti-defense proteins that degrade or sequester oligonucleotides. Because this system only activates when cyclic nucleotides are absent, viral anti-defense proteins that aim to subvert immune responses by depleting host messengers10,11 paradoxically switch defense on.
The authors conclude that the constitutive cyclase activity by mCpol, supported by structural analysis, reveals a novel signaling paradigm in oligonucleotide-based immunity. This work expands on the recent identification of virus-encoded anti-immunity proteins10,11 to show how host systems can counter virus-mediated immune suppression and suggests the possibility of similar protective roles for cyclic oligonucleotides across the tree of life.
Primary Citation
Doherty, E. E.; Adler, B. A.; Yoon, P. H.; Hsieh, K.; Loi, K.; Armbruster, E. G.; Lahiri, A.; Bolling, C. S.; Wilcox, X. E.; Akkati, A.; Iavarone, A. T.; Pogliano, J.; Doudna, J. A. A Miniature CRISPR–Cas10 Enzyme Confers Immunity by Inhibitory Signalling. Nature 2025. https://doi.org/10.1038/s41586-025-09569-9.
References
(1) Wein, T.; Sorek, R. Bacterial Origins of Human Cell-Autonomous Innate Immune Mechanisms. Nat Rev Immunol 2022, 22 (10), 629–638. https://doi.org/10.1038/s41577-022-00705-4.
(2) Doherty, E. E.; Adler, B. A.; Yoon, P. H.; Hsieh, K.; Loi, K.; Armbruster, E. G.; Lahiri, A.; Bolling, C. S.; Wilcox, X. E.; Akkati, A.; Iavarone, A. T.; Pogliano, J.; Doudna, J. A. A Miniature CRISPR–Cas10 Enzyme Confers Immunity by Inhibitory Signalling. Nature 2025. https://doi.org/10.1038/s41586-025-09569-9.
(3) Sullivan, A. E.; Nabhani, A.; Izrailevsky, D. S.; Schinkel, K.; Hoffman, C. R. K.; Robbins, L. K.; Nagy, T. A.; Duncan, M. L.; Ledvina, H. E.; Erbse, A. H.; Kibby, E. M.; Tak, U.; Dinh, D. M.; Ednacot, E. M. Q.; Nguyen, C. M.; Burroughs, A. M.; Aravind, L.; Whiteley, A. T.; Morehouse, B. R. The Panoptes System Uses Decoy Cyclic Nucleotides to Defend against Phage. Nature 2025. https://doi.org/10.1038/s41586-025-09557-z.
(4) Burroughs, A. M.; Zhang, D.; Schäffer, D. E.; Iyer, L. M.; Aravind, L. Comparative Genomic Analyses Reveal a Vast, Novel Network of Nucleotide-Centric Systems in Biological Conflicts, Immunity and Signaling. Nucleic Acids Research 2015, 43 (22), 10633–10654. https://doi.org/10.1093/nar/gkv1267.
(5) Kazlauskiene, M.; Kostiuk, G.; Venclovas, Č.; Tamulaitis, G.; Siksnys, V. A Cyclic Oligonucleotide Signaling Pathway in Type III CRISPR-Cas Systems. Science2017, 357 (6351), 605–609. https://doi.org/10.1126/science.aao0100.
(6) Niewoehner, O.; Garcia-Doval, C.; Rostøl, J. T.; Berk, C.; Schwede, F.; Bigler, L.; Hall, J.; Marraffini, L. A.; Jinek, M. Type III CRISPR–Cas Systems Produce Cyclic Oligoadenylate Second Messengers. Nature 2017, 548 (7669), 543–548. https://doi.org/10.1038/nature23467.
(7) You, L.; Ma, J.; Wang, J.; Artamonova, D.; Wang, M.; Liu, L.; Xiang, H.; Severinov, K.; Zhang, X.; Wang, Y. Structure Studies of the CRISPR-Csm Complex Reveal Mechanism of Co-Transcriptional Interference. Cell 2019, 176 (1–2), 239-253.e16. https://doi.org/10.1016/j.cell.2018.10.052.
(8) Sofos, N.; Feng, M.; Stella, S.; Pape, T.; Fuglsang, A.; Lin, J.; Huang, Q.; Li, Y.; She, Q.; Montoya, G. Structures of the Cmr-β Complex Reveal the Regulation of the Immunity Mechanism of Type III-B CRISPR-Cas. Mol Cell 2020, 79 (5), 741-757.e7. https://doi.org/10.1016/j.molcel.2020.07.008.
(9) Osawa, T.; Inanaga, H.; Numata, T. Crystal Structure of the Cmr2-Cmr3 Subcomplex in the CRISPR-Cas RNA Silencing Effector Complex. J Mol Biol 2013, 425 (20), 3811–3823. https://doi.org/10.1016/j.jmb.2013.03.042.
(10) Nomburg, J.; Doherty, E. E.; Price, N.; Bellieny-Rabelo, D.; Zhu, Y. K.; Doudna, J. A. Birth of Protein Folds and Functions in the Virome. Nature 2024, 633 (8030), 710–717. https://doi.org/10.1038/s41586-024-07809-y.
(11) Hobbs, S. J.; Nomburg, J.; Doudna, J. A.; Kranzusch, P. J. Animal and Bacterial Viruses Share Conserved Mechanisms of Immune Evasion. Cell 2024, 187 (20), 5530-5539.e8. https://doi.org/10.1016/j.cell.2024.07.057.