Computational protein design methods have excelled in the design of static protein structures, but the vast majority of natural proteins switch between two or more conformational states to carry out their functions. In this work, the authors develop a computational workflow, “Conformational Biasing” (CB), for the fast and scalable prediction of mutations that bias proteins toward desired conformational states. By tuning the conformational landscape of proteins, CB can engineer a wide range of protein activities, such as binding, catalysis, and signal transduction. Thus, CB enables mechanistically-informed improvement of conformation-specific functions across diverse protein types.
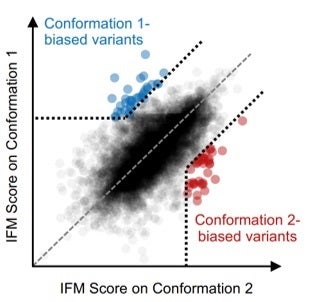
CB uses an inverse folding model (IFM) (1) to generate sequence- and structure-conditioned scores for a library of variants (typically point mutations) of a protein of interest, against two or more alternative backbone structures (Figure 1). Highly divergent scores are used to predict conformation-biasing mutations.
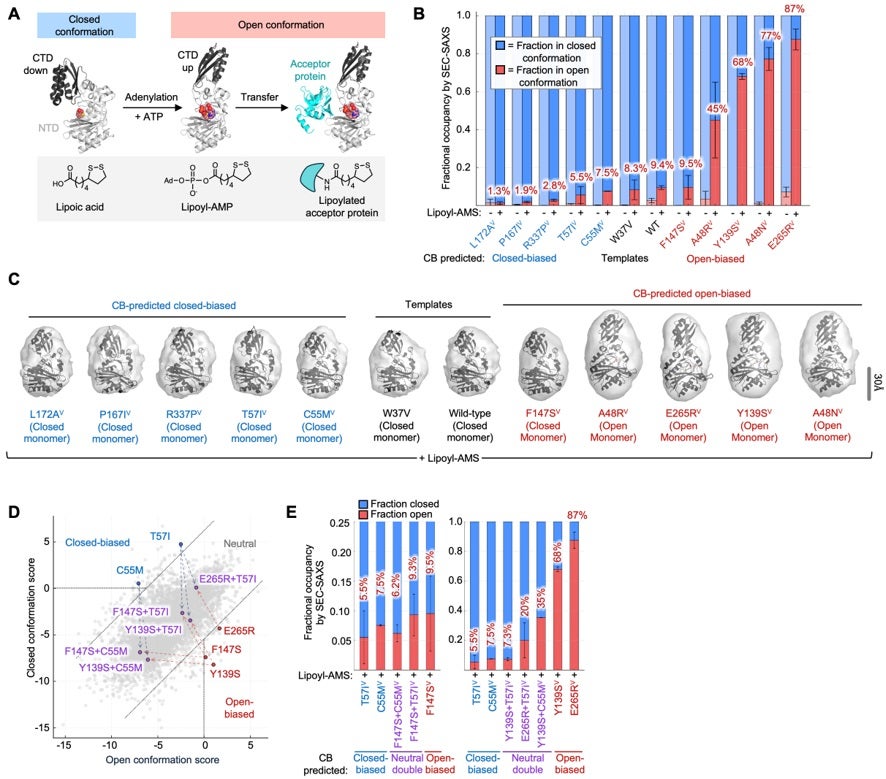
The authors applied CB to the E. coli enzyme Lipoic Acid Ligase A (LplA) both to experimentally evaluate CB’s ability to tune conformational preference and to study the role of conformational changes in LplA’s function. They used size-exclusion chromatography coupled small-angle X-Ray scattering (SEC-SAXS) to directly measure LplA’s size and shape in solution (2). This approach has previously resolved protein conformational ensembles (3) and is suitable for mid-sized proteins like the 38 kDa LplA which may be too large for NMR, and too small for cryo-EM.
The authors developed an automated Oligomer-based analysis method to fit SAXS scattering curves with a modeled mixture of open and closed LplA conformers (4,5). The SAXS data in Figure 2B-C show that four out of five LplA variants predicted by CB to be open-biased displayed a greater occupancy of the open conformation when bound to lipoyl-AMS than W37V template. On the other hand, all five variants predicted to be closed-biased demonstrated a smaller shift to the open state than W37V.
If CB-predicted mutations act on the same conformational equilibrium, then combining an open-biasing mutation and closed-biasing mutation in the same protein should produce an intermediate conformational distribution. To test this, the authors purified five double mutants of LplA, combining both stronger and weaker CB-predicted biasing mutations (Figure 2D). SEC-SAXS measurements showed that the conformational occupancies of these double mutants generally fell in between those of the original single mutants (Figure 2E).
The authors went on to show that conformationally-biased variants of LplA show tuned promiscuous activity levels, enabling the mechanistically-informed design of highly specific and promiscuous LplA variants for labelling applications (6,7)
The resolution and clean disambiguation of monomeric protein that is afforded by SEC-SAXS enabled the deconvolution of conformational states in LplA, and thus the evaluation of the CB methodology. Experiments in this study were performed at the BioSAXS BL4-2 beamline at the Stanford Synchrotron Radiation Lightsource (SSRL).
Primary Citation
P. E. Cavanagh, A. G. Xue, S. A. Dai, A. Qiang, T. Matsui and A. Y. Ting, "Computational Design of Conformation-biasing Mutations to Alter Protein Functions", Science 391, eadv7953 (2026) doi: 10.1126/science.adv7953
References
1. J. Dauparas et al., “Robust Deep Learning-based Protein Sequence Design Using ProteinMPNN”, Science 378, 49-56 (2022).
2. T. Matsui, I. Rajkovic, B. H. M. Mooers, P. Liu, T. M. Weiss, “Adaptable SEC-SAXS Data Collection for Higher Quality Structure Analysis in Solution”, Protein Sci. 33, e4946 (2024).
3. A. H. Phillips et al., “Internal Motions Prime cIAP1 for Rapid Activation”, Biophys. J. 106, 253a (2014).
4. P. V. Konarev, V. V. Volkov, A. V. Sokolova, M. H. J. Koch, D. I. Svergun, “PRIMUS: a Windows PC-based System for Small-angle Scattering Data Analysis”, J. Appl. Crystallogr. 36, 1277-1282 (2003).
5. P. E. Cavanagh, A. G. Xue, S. A. Dai, A. Qiang, T. Matsui and A. Y. Ting, "Computational Design of Conformation-biasing Mutations to Alter Protein Functions", Science 391, eadv7953 (2026) doi: 10.1126/science.adv7953
6. Uttamapinant et al., “A Fluorophore Ligase for Site-specific Protein Labeling Inside Living Cells”, Proc. Natl. Acad. Sci. U. S. A. 107, 10914-10919 (2010).
7. W. Qin, K. Cho, P. E. Cavanagh, A. Ting, “Deciphering Molecular Interactions by Proximity Labeling”, Nat. Methods 18, 133-143 (2021).