|
Life depends on the biochemical activity of the thousands of proteins that
inhabit and decorate the surface of every one of our cells. Proteins
themselves, although simple linear combinations of the twenty amino acids,
derive their remarkable properties from the complex three-dimensional
structures into which they fold. In this way, enzyme active sites are created,
protein-protein recognition surfaces are formed, and the chemistry of life is
set in motion. Although in principle the precise three-dimensional structure
for each protein is encoded in its linear chain of amino acids, in practice it
is often difficult or impossible for a protein to achieve this final fold on
its own in the context of a cellular environment that is packed to the gills
with millions of other proteins, nucleic acids, carbohydrates, lipids, and
other small molecules. As a result, cells have evolved a corps of proteins
known as molecular chaperones that assist newly synthesized proteins as they
adopt their active fold. One such family of chaperones is known as the hsp90
family (Pratt and Toft, 2003). "Client" proteins of the hsp90 family are
diverse, and their functions range from signal transduction to immune response.
Specific inhibitors of hsp90 chaperones exhibit potent anti-tumor activity
(Chiosis et al., 2006; Sharp and Workman, 2006), showing that preventing the
proper folding of client proteins, many of which are implicated in cancer, can
have profound therapeutic implications.
|  |
|
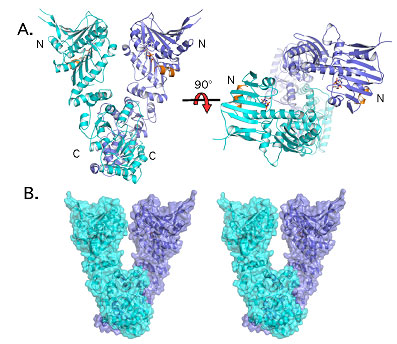
Figure 1. Overview of the GRP94 structure. The two protomers of the GRP94
dimer are shown in blue and cyan. (A) Ribbon drawing of side and top views.
The two N-terminal domains of the dimer do not interact, causing the
misalignment of ATP hydrolysis residues. (B) Stereo surface view of the GRP94
dimer. The twisted V shape is readily apparent.
|
|
The mechanism by which hsp90 chaperones act to mature their client proteins is
not yet established. Hsp90s exist as dimers and it has been shown that
chaperoning activity is closely tied to their ability to hydrolyze ATP
(Obermann et al., 1998; Panaretou et al., 1998; Chadli et al., 2000). In order
to understand how these are related, we used diffraction data collected at
beamlines 11-1 of SSRL and 8.2.1 of ALS to determine the high resolution X-ray
crystal structure of mammalian GRP94, the hsp90 chaperone that is found in the
endoplasmic reticulum of cells. GRP94 is a particularly intriguing member of
the hsp90 family. Earlier studies had suggested that GRP94 did not hydrolyze
ATP, and thus was mechanistically different from other hsp90s (Nicchitta,
1998). The structure that we solved helped explain these observations (Dollins
et al., 2007). In particular, we saw that in the presence of an ATP analog the
GRP94 dimer adopted a structure that resembled a "twisted V" (Figure 1). This
conformation prevented the proper alignment of the residues thought to be
required for ATP hydrolysis. Surprisingly, however, the X-ray structure also
showed that a simple 90 degree rotation of one of the domains of GRP94 could
lead to the productive alignment of the catalytic residues. Prompted by this
structural insight, we carried out a series of careful biochemical measurements
that showed that in fact GRP94 had a very weak but reproducible ATPase
activity. These experiments suggested that the transition from the "twisted V"
conformation to one that aligns the catalytic residues was likely to be a key
step in the regulation of GRP94 activity (Figure 2). This insight was
important not only for our understanding of GRP94 but also for understanding
other hsp90s. In particular, unlike its counterparts in yeast or bacteria,
cytoplasmic human Hsp90 also exhibits unusually weak ATPase activity, and thus
may bear a strong structural resemblance to GRP94. Together these observations
have succeeded in establishing the place of GRP94 in the hsp90 family and,
together with our earlier studies of the isolated domains of GRP94 (Soldano et
al., 2003; Immormino et al., 2004; Dollins et al., 2005), opens the door to the
design of inhibitors that specifically target this chaperone.
|  | |
|
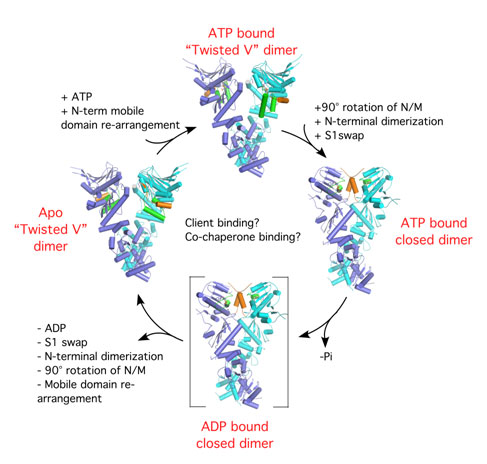
Figure 2. Model of the GRP94 ATP hydrolysis mechanism. The conformational
changes that lead to the alignment of ATP-catalytic residues are shown. Such
rearrangements are likely to allow for the binding and release of client
proteins from the chaperone.
| |
|
Primary Citation
Dollins, D.E., Warren, J.J., Immormino, R.M., and Gewirth, D.T. (2007).
Structures of GRP94-nucleotide complexes reveal mechanistic differences between
the hsp90 chaperones. Mol Cell 28, 41-56.
References
|
Chadli, A., Bouhouche, I., Sullivan, W., Stensgard, B., McMahon, N., Catelli,
M.G., and Toft, D.O. (2000). Dimerization and N-terminal domain proximity
underlie the function of the molecular chaperone heat shock protein 90. Proc
Natl Acad Sci U S A 97, 12524-12529.
Chiosis, G., Rodina, A., and Moulick, K. (2006). Emerging Hsp90 inhibitors:
from discovery to clinic. Anticancer Agents Med Chem 6, 1-8.
Dollins, D.E., Immormino, R.M., and Gewirth, D.T. (2005). Structure of
Unliganded GRP94, the Endoplasmic Reticulum Hsp90: Basis for Nucleotide-Induced
Conformational Change. J Biol Chem 280, 30438-30447.
Dollins, D.E., Warren, J.J., Immormino, R.M., and Gewirth, D.T. (2007).
Structures of GRP94-nucleotide complexes reveal mechanistic differences between
the hsp90 chaperones. Mol Cell 28, 41-56.
Immormino, R.M., Dollins, D.E., Shaffer, P.L., Soldano, K.L., Walker, M.A., and
Gewirth, D.T. (2004). Ligand-induced conformational shift in the N-terminal
domain of GRP94, an Hsp90 chaperone. J Biol Chem 279, 46162-46171.
Nicchitta, C.V. (1998). Biochemical, cell biological and immunological issues
surrounding the endoplasmic reticulum chaperone GRP94/gp96. Curr Opin Immunol
10, 103-109.
Obermann, W.M., Sondermann, H., Russo, A.A., Pavletich, N.P., and Hartl, F.U.
(1998). In vivo function of Hsp90 is dependent on ATP binding and ATP
hydrolysis. J Cell Biol 143, 901-910.
Panaretou, B., Prodromou, C., Roe, S.M., O'Brien, R., Ladbury, J.E., Piper,
P.W., and Pearl, L.H. (1998). ATP binding and hydrolysis are essential to the
function of the Hsp90 molecular chaperone in vivo. Embo J 17,
4829-4836.
Pratt, W.B., and Toft, D.O. (2003). Regulation of signaling protein function
and trafficking by the hsp90/hsp70-based chaperone machinery. Exp Biol Med
(Maywood) 228, 111-133.
Sharp, S., and Workman, P. (2006). Inhibitors of the HSP90 molecular chaperone:
current status. Adv Cancer Res 95, 323-348.
Soldano, K.L., Jivan, A., Nicchitta, C.V., and Gewirth, D.T. (2003). Structure
of the N-terminal domain of GRP94. Basis for ligand specificity and regulation.
J Biol Chem 278, 48330-48338.
|
| PDF version | | Lay Summary | |
Highlights Archive
|
| SSRL is supported
by the Department of Energy, Office of Basic Energy Sciences. The SSRL
Structural Molecular Biology Program is supported by the Department of Energy,
Office of Biological and Environmental Research, and by the National Institutes
of Health, National Center for Research Resources, Biomedical Technology
Program, and the National Institute of General Medical Sciences. |
|



