
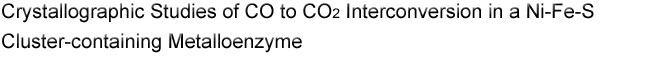
While scientists and engineers struggle to develop technologies to remove CO
and CO2 from our environment, anaerobic bacteria such as
Rhodospirillum rubrum and Carboxydothermus hydrogenoformans have
the ability to utilize the gaseous pollutant CO as their sole carbon and energy
source (1-2). This ability derives from the
reversible oxidation of CO to CO2 catalyzed at a Ni-Fe-S active site
metal cluster (C-cluster) of the enzyme carbon monoxide dehydrogenase (CODH).
Acetogenic bacteria such as Moorella thermoacetica also use CODH in a
bifunctional CODH/acetyl-CoA synthase (ACS) enzyme complex to first convert the
greenhouse gas CO2 to a CO intermediate. CO then travels through a
tunnel within the CODH/ACS complex to the ACS subunit's active site
metallocluster (A-cluster) where it is combined with a methyl group and
coenzyme A to form acetyl-CoA. Acetoclastic methanogens also harbor CODH and
ACS subunits in the enzyme complex acetyl-CoA decarbonylase/synthase (ACDS);
however, in this case, the reaction catalyzed is the degradation of acetyl-CoA
to form CO2 and another greenhouse gas, methane (CH4). At
the heart of these reaction pathways that are vital to the global carbon cycle
is the reversible oxidation of CO to CO2 catalyzed by the active
site C-cluster of CODH. In this work, we have used X-ray crystallography
performed at SSRL (Beam Line 11-1) and at ALS (Beam Line 5.0.1) to understand
the chemistry of this remarkable metallocluster.
Figure 1.
Representative structures of CODH C-clusters (a) with and (b)
without the bridging sulfide. The C-cluster shown in (a) is from the structure
of C. hydrogenoformans CODH (ChCODH) (PDB ID: 1SU8) and is shown
in cyan cartoon with residue numbering following the ChCODH sequence.
The bridging sulfide is circled in red. The C-cluster in (b) is from the
structure of M. thermoacetica CODH/ACS (MtCODH/ACS) solved
previously (PDB ID: 1MJG) and is shown in gray cartoon with residue numbering
following the MtCODH/ACS sequence. C-clusters are depicted in
ball-and-stick, while ligands are shown as sticks: Ni in green, Fe in orange, S
in yellow, and N in blue. Ni and the unique Fe of the C-cluster are labeled as
Ni and Fe, respectively.
In this study, we have solved two crystal structures of the
M. thermoacetica
CODH/ACS complex, with the C-cluster bound by substrate and inhibitor
molecules. The first is a native structure, illustrating a substrate water
molecule bound to the unique Fe of the C-cluster (Figure 2a). After soaking
native crystals in a solution containing potassium cyanide, we obtained a
second crystal structure where the substrate water molecule is still seen bound
to the unique Fe, but cyanide, an inhibitor which mimics CO binding, is bound
to Ni, adjacent the substrate water molecule (Figure 2B). Importantly, neither
structure contains the sulfide bridge. With the substrate binding sites now
identified, we have contributed in both determining of the catalytic relevance
of the sulfide bridge and in uncovering the mechanism of the C-cluster. From
the results of this and other recent crystallographic studies
(9-10), we have
reached a unified catalytic mechanism of the C-cluster that excludes the
bridging sulfide. Given the central role of CODH in the global carbon cycle,
our detailed mechanistic understanding of this important enzyme may have
broader applications in biotechnology and environmental health.
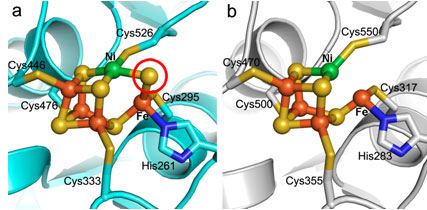
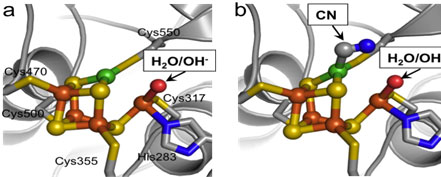
Figure 2.
The (a) native and (b) cyanide-bound C-clusters from
MtCODH/ACS, using the same
representation and coloring as in Figure 1b.
Primary Citation
Kung, Y., Doukov, T.I., Seravalli, J., Ragsdale, S.W., Drennan, C.L. (2009)
Crystallographic snapshots of cyanide- and water-bound C-clusters from
bifunctional carbon monoxide dehydrogenase/acetyl-CoA synthase.
Biochemistry. 48, 7432-7440.
References
The C-cluster is a Ni-Fe-S cluster whose metal content and geometry are
unprecedented in biology. Here, Ni is located within a distorted cubane, along
with three Fe and four S atoms, with a unique Fe coordinated nearby, liganded
by a histidine residue. While several previous crystallographic studies
confirmed this distinct arrangement of metals, there was conflicting structural
data on the presence (3-4) or absence
(5-8) of an additional sulfide ligand
bridging the Ni and unique Fe (Figure 1A-B). As the sulfide bridge occupies
putative substrate binding sites, its presence or absence is of considerable
catalytic significance. Because these CODH structures did not contain any
substrates or analogs bound to the C-cluster, the role of the bridging sulfide
and the mechanism of the cluster had been unclear and hotly debated.
SSRL is supported by the Department of Energy, Office of Basic Energy Sciences. The SSRL Structural Molecular Biology Program is supported by the Department of Energy, Office of Biological and Environmental Research, and by the National Institutes of Health, National Center for Research Resources, Biomedical Technology Program, and the National Institute of General Medical Sciences.