
 Andrey S. Krasilnikov*, Xiaojing Yang*, Tao Pan†, and Alfonso
Mondragón*‡
Andrey S. Krasilnikov*, Xiaojing Yang*, Tao Pan†, and Alfonso
Mondragón*‡
*Department of Biochemistry, Molecular Biology and Cell Biology, Northwestern University, Evanston, IL 60208, USA
†Department of Biochemistry and Molecular Biology, University of Chicago, 920 E. 58th St., Chicago, IL 60637 USA
Transfer RNA (tRNA) is produced as a precursor molecule that needs to be processed both at its 3' and 5' ends. Ribonuclease P, or RNase P, is the only endonuclease responsible for processing the 5' end of tRNA by cleaving a precursor and leading to tRNA maturation. It contains an RNA and a protein component and has been identified in all organisms. It is one of the first catalytic RNA molecules (ribozymes) identified and its study has been pivotal to our understanding of the role of RNA molecules in catalysis. RNase P is a true multi-turnover ribozyme and one of only two ribozymes, together with the ribosome, conserved in all three kingdoms of life (for reviews, see (Altman and Kirsebom, 1999; Frank and Pace, 1998)).
The bacterial RNase P RNA component (or P RNA) consists of between 350-450 nucleotides, while the protein component is a small, basic protein of around 120 amino acids. In a manner similar to proteins, large RNA molecules can be formed by different domains that fold independently. Eubacterial P RNA consists of two domains: a specificity domain or S-domain and a catalytic domain or C-domain (Loria and Pan, 1996; Pan, 1995). The S-domain alone can bind pre-tRNA directly (Qin et al., 2001).
| ||
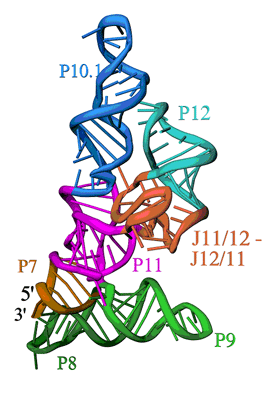
| Figure 1. Ribbon diagram of the S-domain of B. subtilis RNase P. The P7-P11 stems form a cruciform, with one helix in the cruciform formed by the stacking of P8 and P9, and the other by the stacking of P7, P10 and P11. The GAAA tetraloop at the end of P12 interacts with the tetraloop receptor in P10.1. The highly conserved J12/11-J11/12 module forms a large structure devoid of Watson-Crick base pairs. | ||
In our work, we report the crystal structure of the 154 nucleotide long S-domain of Bacillus subtilis RNase P at 3.15 Ĺ resolution (Krasilnikov et al., 2003). This is the second largest atomic resolution structure of an RNA molecule solved so far that is not in complex with proteins. Solving this structure involved paying meticulous attention to all aspects of the structure elucidation process, from the way the crystals were grown and handled, to the way data were processed and analyzed. The S-domain crystals are very sensitive to any environmental changes, and hence it was crucial to develop a protocol for handling and cryo-cooling the crystals that preserved isomorphism among them. Additionally, as the crystals diffracted very weakly, all data collection was done at a synchrotron source. In order to solve the structure, we had to screen for heavy atom derivatives as we could not easily incorporate an anomalous scatterer into the structure. Although the final data sets were collected at DND CAT at the Advanced Photon Source, much of the necessary preliminary work was done at SSRL using Beam Line 9-1. In particular, the search for optimal handling conditions and ways of preserving isomorphism were prompted to a large extent by the results we obtained during our visits to SSRL. Many crystals had to be screened and analyzed and access to SSRL was very important for the project.
The overall structure is shown in Figure 1. The S-domain consists of several distinct secondary structure modules, which were predicted from the sequence of the P RNA and phylogenetic comparison. The most relevant features are: a cruciform formed by the stacked P7, P10 and P11 and the stacked P8 and P9 helices, b) the packing of the P10.1 and P12 helices through a GAAA tetraloop-tetraloop receptor interaction, and c) an unusually folded module linking P11 and P12 (J11/12-J12/11), which contains a large number of universally conserved nucleotides, and is stabilized without canonical Watson-Crick base pairing. While the S-domain forms a well-packed and compact structure, it is important to note that the P11 and P9 helices together with the J11/12-J12/11 module form a clamp-like opening that contains nucleotides involved in pre-tRNA binding and is large enough to accommodate the TyC stem-loop of a pre-tRNA molecule.
The structure allowed us to make important observations regarding the general
architecture of the S-domain, the structure of a region conserved in RNase P's
of all organisms and the position of nucleotides involved in interactions with
the substrate, and in this way defining regions that are crucial for activity.
Additionally, the structure of the S-domain of B. subtilis RNase P
provides a molecular framework for studying many of the interactions that must
occur during tRNA processing by this crucial ribozyme. The structure is
consistent with all the available data for bacterial RNase P's and extends our
understanding of RNase P across all kingdoms of life. Furthermore, as an
important addition to the still limited number of large RNA's of known
structure, it advances our knowledge of the general principles of RNA structure
and packing.
References
- Altman, S., and Kirsebom, L. A. (1999). In The RNA World, R. F. Gesteland, T. R. Cech, and J. F. Atkins, eds. (Cold Spring Harbor, New York, Cold Spring Harbor Laboratory Press), pp. 351-380.
- Frank, D. N., and Pace, N. R. (1998). Ribonuclease P: unity and diversity in a tRNA processing ribozyme. Annu Rev Biochem 67, 153-180.
- Loria, A., and Pan, T. (1996). Domain structure of the ribozyme from eubacterial ribonuclease P. RNA 2, 551-563.
- Pan, T. (1995). Higher order folding and domain analysis of the ribozyme from Bacillus subtilis ribonuclease P. Biochemistry 34, 902-909.
- Qin, H., Sosnick, T. R., and Pan, T. (2001). Modular construction of a tertiary RNA structure: the specificity domain of the Bacillus subtilis RNase P RNA. Biochemistry 40, 11202-11210.
- Krasilnikov, A. S., Yang, X., Pan, T., and Mondragón, A. (2003). Crystal structure of the specificity domain of ribonuclease P. Nature 421 760-764.
SSRL Highlights Archive