
 Membrane proteins are notoriously difficult to crystallize, and fiber-forming
proteins were actually declared "uncrystallizable" by the eminent x-ray
crystallographer Sir Lawrence Bragg. Supported by the facilities and staff at
SSRL, a team of researchers has recently determined structures that solved both
problems by defining the atomic structure of the membrane-associate protein
pilin and its assembled Type IV pilus (T4P) fiber structure. T4P are
filamentous organelles displayed on the surfaces of most Gram-negative bacteria
1. T4P are central to host colonization for many bacterial pathogens, mediating
diverse and essential functions such as motility, adhesion, microcolony
formation and uptake of DNA and specific filamentous phage. T4P are several
microns in length and only 60-90 Ĺ in diameter, and are comprised of thousands
of copies of a single subunit, the pilin protein. Type IV pilins from different
bacterial species share a common sequence of mostly hydrophobic amino acids in
their N-terminus, as well as a pair of cysteines in their C-terminus, but
differ substantially beyond these sites. Their prominent exposure on the
bacterial surface and their key functions in virulence make T4P attractive
targets for vaccines and therapeutics, the design of which would greatly
benefit from their detailed molecular structures. Structural analyses of the
Type IV pili have, however, proved to be extremely challenging. The presence of
the hydrophobic N-terminal segment, which allows pilin to assemble and
disassemble from the cell membranes, makes the pilin subunits insoluble without
detergent. This problem has hindered efforts to crystallize the full-length
proteins. Furthermore, the extremely thin and featureless pilus filaments
reveal scant information on their helical symmetry, and thus resist classical
helical image reconstruction approaches using electron micrograph (EM) images.
Membrane proteins are notoriously difficult to crystallize, and fiber-forming
proteins were actually declared "uncrystallizable" by the eminent x-ray
crystallographer Sir Lawrence Bragg. Supported by the facilities and staff at
SSRL, a team of researchers has recently determined structures that solved both
problems by defining the atomic structure of the membrane-associate protein
pilin and its assembled Type IV pilus (T4P) fiber structure. T4P are
filamentous organelles displayed on the surfaces of most Gram-negative bacteria
1. T4P are central to host colonization for many bacterial pathogens, mediating
diverse and essential functions such as motility, adhesion, microcolony
formation and uptake of DNA and specific filamentous phage. T4P are several
microns in length and only 60-90 Ĺ in diameter, and are comprised of thousands
of copies of a single subunit, the pilin protein. Type IV pilins from different
bacterial species share a common sequence of mostly hydrophobic amino acids in
their N-terminus, as well as a pair of cysteines in their C-terminus, but
differ substantially beyond these sites. Their prominent exposure on the
bacterial surface and their key functions in virulence make T4P attractive
targets for vaccines and therapeutics, the design of which would greatly
benefit from their detailed molecular structures. Structural analyses of the
Type IV pili have, however, proved to be extremely challenging. The presence of
the hydrophobic N-terminal segment, which allows pilin to assemble and
disassemble from the cell membranes, makes the pilin subunits insoluble without
detergent. This problem has hindered efforts to crystallize the full-length
proteins. Furthermore, the extremely thin and featureless pilus filaments
reveal scant information on their helical symmetry, and thus resist classical
helical image reconstruction approaches using electron micrograph (EM) images.
John Tainer and colleagues at the Scripps Research Institute have succeeded in
solving the crystal structures of Type IV pilins from several important human
pathogens using the macromolecular crystallography beamlines at SSRL (Beamlines
7-1, 9-1, 9-2 and 11-1). This success built directly upon the earlier efforts
of Tainer group members Hans Parge and Andrew Arvai who made the original
breakthrough pilin structure determination as aided enormously by the then new
area detectors installed at SSRL according to John Tainer 2. Full length
pilin structures were determined for Neisseria gonorrhoeae and
Pseudomonas aeruginosa, and a truncated structure lacking the
hydrophobic N-terminus was determined for
Vibrio cholerae 2, 3. In a recently published article in
Molecular Cell, this
group report the 2.3 Ĺ structure of N. gonorrhoeae (GC) pilin along with
a "pseudo-atomic" structure of the pilus filament, solved by combined cryoEM
reconstruction methods and computational docking of the pilin crystal
structure 4. The N. gonorrhoeae Type IV pili are
displayed peritrichously on the bacterial surface and mediate a multitude of
functions in virulence, including immune escape. They share substantial
sequence identity with the Type IV pili of N. meningitidis, which causes
meningitis. The GC pilin crystal structure nicely extends the previous
structure solved by the same group 2, and furthermore reveals
the precise identity of two unusual post-translational modifications: an
O-linked galactose (a1→3)
diacetamidodideoxyglucose at Ser63 and a phosphoethanolamine at Ser68 (Fig.
1A). These post-translational modifications undergo phase variation and may
also vary in structure from one generation of N. gonorrhoeae to the
next, and have been proposed to function in immune escape. Another unusual
feature of the pilin subunits is the "hypervariable loop", a segment between
residues 128 and 141 that undergoes extreme sequence variation. This loop
protrudes from the back of the pilin globular domain. The remainder of the GC
pilin protein looks similar to other pilins: the hydrophobic N-terminus forms
an extended a-helix, half of which is embedded in a
b-sheet within the C-terminal globular domain.
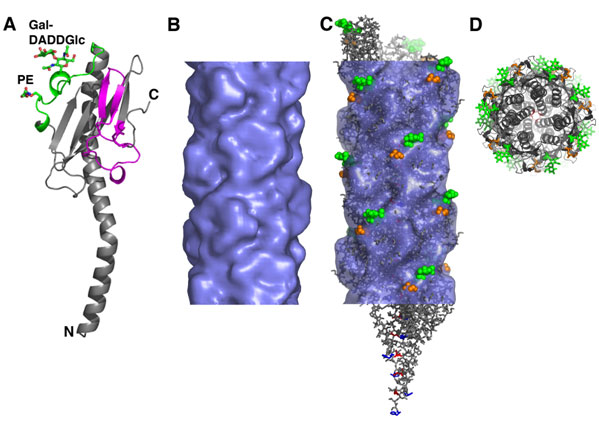
Figure 1 Structure of the N. gonorrhoeae Type IV pilus by
x-ray crystallography and cryoEM reconstruction. (A) GC pilin structure at
2.3 Ĺ showing the postranslational modifications galactose (a1→3) diacetamidodideoxyglucose (Gal-DADDGlc) at Ser63
and a phosphoethanolamine (PE) at Ser68. The hypervariable loop is colored
magenta. (B) CryoEM reconstruction of the GC pilus filament and
(C) the pseudoatomic resolution structure of the filament fit into the
EM density. (D) End view of the pilus structure showing the packing of
the N-terminal a-helices and the protruding
postranslational modifications and hypervariable loop.
Tainer and colleagues collaborated with Ed Egelman at the University of
Virginia to determine a cryoEM reconstruction of the GC pilus filament using
Egelman's Iterative Helical Real Space Reconstruction software 5. Although the
filaments appear to be very smooth and featureless when examined by EM, they
are in fact highly corrugated (Fig. 1B). The pilus structure was built by
computationally fitting the pilin subunit structure into the cryoEM density and
generating a filament using the symmetry operators determined for the
reconstruction. In this pseudoatomic resolution pilus structure, deep grooves
run between the subunits, and ridges formed by the post-translationally
modified regions and the hypervariable loop protrude from the filament surface
(Fig. 1C). The filaments are held together primarily by the N-terminal
segments, which form a helical array in the core of the filament (Fig. 1D).
The N. gonorrhoeae Type IV pilus structure provides important insights
into both pilus assembly and functions in pathogenesis. The filament can be
viewed as three helical pilin polymers that twist around each other. These
polymers are likely to be assembled simultaneously at the inner membrane of the
bacteria, from a reservoir of pilin subunits that are anchored in the membrane
via their hydrophobic N-termini. Importantly, the pilin and pilus structures
provided the basis for a unified model for the assembly of these filaments that
was recently supported by structures of the secretion superfamily ATPase
6,
which were also done partly at SSRL and show the basis for the piston-like
shifts to push T4P out by two helical turns. The grooves that run along the
filament surface are positively-charged and represent an ideal, non-specific
binding site for DNA, which can be brought into the cell by pilus retraction.
And the locations of the variable post-translational modifications and
hypervariable loop on protruding ridges on the filament would mask recognition
of more conserved regions of the protein by protective antibodies. This
structure is the first such structure determined for a Type IV pilus filament.
The overall architecture of the filament is expected to be shared by all Type
IV pili based on sequence and structure conservation for the pilin subunits. It
is the regions of the subunit that vary, quite dramatically, among the pilins,
that are exposed on the pilus surface and define its chemistry and hence its
functions. This molecular understanding of a Type IV pilus filament reveals new
targets for antibacterial agents that may have broad specificity.
This work was supported by NIH grants AI22160 (JAT, LC), EB001567 (EHE) and
GM076503 (NV) and a fellowship from The Canadian Institutes of Health Research
(LC).
Primary Citation
References
Craig, L. et al. Type IV Pilus Structure by Cryo-Electron Microscopy and
Crystallography: Implications for Pilus Assembly and Functions. Molecular Cell
23, 651-662 (2006).
| SSRL is supported by the Department of Energy, Office of Basic Energy Sciences. The SSRL Structural Molecular Biology Program is supported by the Department of Energy, Office of Biological and Environmental Research, and by the National Institutes of Health, National Center for Research Resources, Biomedical Technology Program, and the National Institute of General Medical Sciences. |