
Copper is a
required micronutrient for all living cells, being an essential component of
many metalloenzymes, but free intracellular copper is highly toxic. Because of
this copper within cells is very tightly controlled, with specific copper ion
pumps (both importers and exporters) located at the cell surface, which are
coupled to highly specific metallochaperones that act as molecular taxi-cabs,
transporting copper to the active sites of particular target proteins [1]. The expression of the copper homeostatic apparatus is also
very tightly controlled, and is very sensitive to copper levels. In bacteria,
two families of copper-responsive transcriptional
repressors have been described; these are typified by the CueR repressor in
 | |
Figure 1: Mycobacterium tuberculosis
| |
Escherichia coli [2], and the CopY repressor in
Enterococcus hirae [3].
However, many bacteria, including important human pathogens, such as
Mycobacterium tuberculosis (Figure 1) and Listeria monocytogenes,
contain
neither family, and the mechanism of copper regulation in these organisms has
been unknown. The discovery of the CsoR repressor from M. tuberculosis
represents the first example of a totally new family of bacterial repressors
that appears to be more widespread than either of the two established repressor
families.
|  | |
|
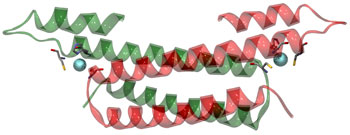
Figure 2: Crystal structure of the CsoR homodimer. The metallic atoms
indicate cuprous ions. [large view]
| | |
In the presence of copper the CsoR repressor dissociates from the cso operon
which can then be expressed. The operon contains the gene for what is believed
to be a cellular copper exporter - CtpV [4]. Binding of Cu(I)
to the CsoR dimer is thought to cause a conformational change, [effecting
decreased] CsoR affinity for DNA, allowing expression of the cso operon, thus
providing a protective mechanism against toxic levels of copper within cells.
The binding of copper to CsoR is exceptionally strong (KCu ≥ 10-19M), which is expected for a system that is
critical for removal of toxic copper from cells. While CsoR binds two Cu(I) per
homodimer only one copper is required to dissociate it from DNA. Liu et
al used a series of techniques to understand the structure of the Cu(I)
bound CsoR repressor. Protein crystallography of Cu(I)-bound CsoR indicates
that CsoR is an alpha-helical dimer, with each protomer composed of three
helices (Figure 2). The copper is bound between the two subunits, coordinated
by the side chains of amino acids from each subunit (Cys36, Cys65˙, and
His61˙). Copper K-edge X-ray absorption spectroscopy (XAS) was used to provide
accurate bond-length information on the copper site of Cu(I)-bound CsoR. XAS
experiments were conducted on SSRL's structural molecular biology XAS beamline
9-3, with additional (preliminary) data being measured on the Canadian Light
Source HXMA beamline. Analysis of the extended X-ray absorption fine structure
(EXAFS) oscillations indicated a three-coordinate site with two Cu-S ligands at
2.21 Ĺ and one oxygen or nitrogen at 2.06 Ĺ. Analysis of the Cu XAS of the
Cu(I)-CsoR H61A mutant indicated a two-coordinate site with two sulfurs at 2.14
Ĺ (Figure 3).
|  | |
|
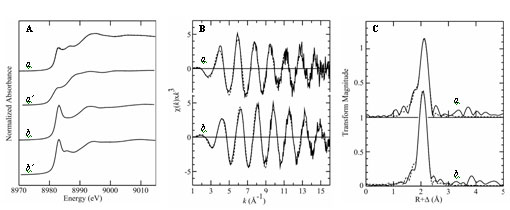
Figure 3: XAS spectra of wild-type CsoR (a) and H61A mutant CsoR (b). A
shows the near-edge spectra compared with those of three-coordinate models (a'
and b', respectively), B shows the EXAFS oscillations and C shows the EXAFS
Fourier transforms (Cu-S phase-corrected). In B and C the solid lines show
experimental data, while the broken lines show the best-fit.
| | | |
Sequence comparisons indicate that homologues of
CsoR are very widespread in prokaryotes, being present in all major classes of
eubacteria. The study of Liu
et al thus represents a structural and functional characterization of the first
known member of a new and widely-distributed family of prokaryotic copper
regulators. The presence of CsoR is likely to enhance the survival of
M. tuberculosis in the host, and whether CsoR can be exploited as the basis for
new treatments for tuberculosis awaits future research.
Primary Citation
Liu, T., Ramesh, A., Ma, Z., Ward, S. K., Zhang, L., George, G. N., Talaat, A.
M., Sacchettini, G. C., Giedroc, D. P. "CsoR is a novel Mycobacterium
tuberculosis copper-sensing transcriptional regulator". Nature Chem.
Bio., 2007, 3, 60-68
References
-
Rae, T.D., Schmidt, P.J., Pufahl, R.A., Culotta, V.C. & O'Halloran,
T.V. Science 1999, 284, 805-808.
-
Strausak, D. & Solioz, M. J. Biol. Chem. 1997, 272, 8932-8936.
-
Stoyanov, J.V., Hobman, J.L. & Brown, N.L. Mol. Microbiol. 2001,
39,502-511.
-
Palmgren, M.G. & Axelsen, K.B. Biochim. Biophys. Acta 1998, 1365,
37-45
|
| PDF
Version | | Lay Summary | |
Highlights Archive
|
| SSRL is supported
by the Department of Energy, Office of Basic Energy Sciences. The SSRL
Structural Molecular Biology Program is supported by the Department of Energy,
Office of Biological and Environmental Research, and by the National Institutes
of Health, National Center for Research Resources, Biomedical Technology
Program, and the National Institute of General Medical Sciences. |
|




