
by Thiang Yian Wong, Robert Schwarzenbacher and Robert C. Liddington
The Burnham Institute, 10901 North Torrey Pines Road, La Jolla, CA 92037.
Anthrax Toxin is a major virulence factor in the infectious disease,
Anthrax1. This toxin is produced by Bacillus
anthracis,
which is an
encapsulated, spore-forming, rod-shaped bacterium. Inhalation anthrax,
the most deadly form, is contracted through breathing spores. Once
spores germinate within cells of the immune system
called macrophages2,
bacterial cells are released into the bloodstream. There they proliferate
rapidly and secrete Anthrax Toxin, ultimately leading to septic
shock and death. Although antibiotics may be used to kill the bacteria,
the level of toxin has often become so high in the bloodstream that
removing the bacteria alone is not sufficient to prevent death.
Therefore, the design of anti-toxins offers the prospect of treatment in
the advanced stages of infection. Together with collaborators from the
NIH and Harvard Medical School, we are involved in the atomic resolution
study of the Anthrax Toxin components and their complexes, including
small molecules with therapeutic potential. Data collection at SSRL
and other synchrotron radiation sources has been key to the advances made in
this research so far and is expected to play a continuing role in the future.
Anthrax Toxin, secreted into the bloodstream, is composed of 3 individual
proteins - Protective Antigen (PA), Lethal Factor (LF) and Edema Factor
(EF),
which act in concert to disrupt cellular signaling systems in the
host macrophage (Figure 1).
PA binds a receptor on the host cell
surface, is cleaved and activated by a host protease to form PA63,
and then forms a seven-membered ring structure that binds
the two toxic
enzymes, LF and EF1, 3,4. The resulting toxin complex is engulfed
(endocytosed) into intracellular compartments called endosomes. Natural
cellular processes lead to acidification of the endosome, which triggers
a conformational change in PA63, causing it to insert into the
endosomal
membrane and translocates the toxic enzymes into the host cell interior
(cytosol). Once inside, LF and EF do their damage to the host cell
defense system. LF is a metalloproteinase that cleaves six members of
the MAPKK family5 of intracellular signaling
proteins, removing the
specific fragment from individual MAPKKs that is crucial for immediate
interaction with other signaling proteins. This action by LF rapidly
blocks the signals that would normally recruit other immune cells to
fight the infection6. EF, on the other hand, is a
calmodulin-activated adenylyl cyclase that increases the concentration of a
messenger molecule
(cAMP) needed for regulated cell functions7 to abnormal levels, causing
accumulation of fluids within and between cells, and hence edema. The
disruption of normal signaling pathways result in cell lysis, the sudden
release of messenger molecules, and toxic shock.
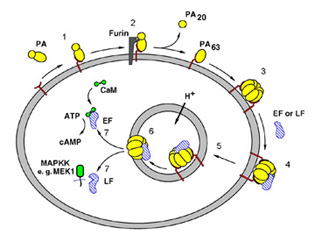
Figure 1. Steps in anthrax intoxication.
1, PA binds to the Anthrax Toxin Receptor3;
2, furin cleaves and releases PA20;
3, PA63 forms a heptamer4; 4, the toxic enzymes bind to
PA63; 5,
receptor-mediated endocytosis; 6, acidification of the endosome leads to
membrane insertion of PA63; 7, translocation of the toxic enzymes
into the cytosol. LF, Lethal Factor; EF, Edema Factor.
The three-dimensional atomic resolution structures of all three
individual components (PA4,
LF8, and EF9)
have now been solved by X-ray crystallography.
We recently reported the crystal structure of Lethal
Factor and its complex with a 16-amino acid residues long (16-mer) peptide
representing the N-terminus
of its natural substrate, MAPKK-2, in Nature (23 October online pre-
release and 8 November 2001 in print)8. Two
different crystal
forms grew out of crystallization conditions: monoclinic P21 and
cubic I4132. The
monoclinic crystal form was used to solve the structure of wild-type
native LF, with zinc bound in the active site, to 2.2 Å resolution.
The cubic crystal form (Figure 2) was instrumental in the introduction of the
target peptide into the active site of LF through peptide solution soaks,
thus leading to the crystal structure of the LF-MAPKK2 N-terminus peptide
complex, to 3.9 Å resolution. All of the data for the cubic LF crystals
and peptide complex were collected at the Stanford Synchrotron Radiation
Laboratory (SSRL), mainly on Beamlines 7-1 and 9-1. Due to the large unit
cell and weak diffraction properties of this crystal form, the use of the
SSRL synchrotron source was essential to obtaining high-resolution data.
Data were collected at SRS, Daresbury, ESRF, Grenoble, APS, Chicago and
National Synchrotron Light Source, Brookhaven to solve the atomic resolution
structures of PA, LF and EF.

Figure 2. An
I4132
space group cubic crystal of the Anthrax Toxin Lethal Factor.
 |
|
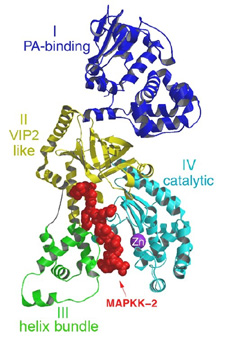
| Figure 3. A ribbon representation of Lethal Factor, colored by domain, with the N-terminus 16mer of MAPKK-2, in red colored ball-and-stick, bound in the active site. This figure was generated using the program MOLSCRIPT, RENDER and RASTER3D10-12. | |
Lethal Factor (LF) comprises 4 structural domains: Domain I provide the binding surface for PA. Domain II resembles a toxin enzyme, VIP2, from Bacillus cereus, although it does not share its catalytic activity. Domain III is inserted into domain II, and has arisen from the duplication of a structural element from domain II. Domain IV, which resembles Domain I, is the catalytic center, with its protein fold resembling the protein family of the zinc-dependent proteinase typified by Thermolysin. Domains II, III and IV together create a groove - 40 Å long - into which a 16 -amino acid long peptide representing the N- terminus of its natural substrate MAPKK-2 binds (Figures 3 and 4). Thus, LF is an interesting example in protein evolution, in which the functional unit arises from gene duplication and fusion of domain building blocks. The structure reveals, for the first time, a protease bound to its uncleaved substrate. The shape of the active site groove allows for specificity but yet gives the freedom for variable target processing. The binding groove in LF is acidic in nature 8, and nicely complements the basic nature of the N-termini of the six MAPKKs recognized by LF5, although higher resolution complexes will be necessary to explore these interactions in atomic detail. Because LF is able to cripple normal host cell functions through several targets all at once, its contributions to the seriousness of Anthrax infections is tremendous, and the key to LF efficiency lies primarily in the active center. Thus, understanding the crystal structure of LF, and atomic interactions with natural targets in its active center, will be crucial in the design of anti-toxins that inhibit LF activity.
 |
| Figure 4. Molecular surface representation of Lethal Factor, colored by charge (red=negative, blue=positive), with the model of the MAPKK-2 N-terminal peptide as ball-and-stick. The electron density surrounding the peptide is a "gradient omit map" calculated at 3.9 Å resolution. Where the protein surface hides the peptide model or map, it is rendered translucent. This figure was prepared using SPOCK13. |
References
SSRL Highlights Archive
