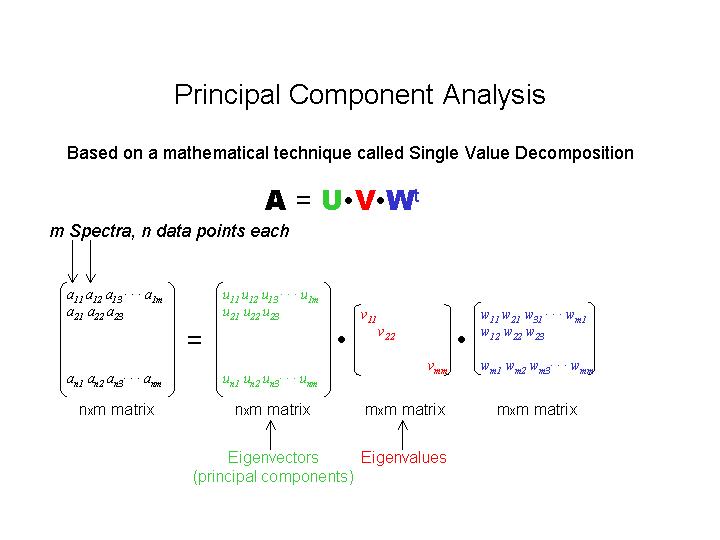



To start the PCA analysis, type:
PCA SAMP.HLDSAMP.HLD is an ascii file containing the m filenames of the spectra of related unknowns. This can be created using a basic text editor, such as NOTEPAD. For example, SAMP.HLD might look like this:
SAMP1.EDGAfter starting PCA, the following appears:
SAMP2.EDG
SAMP3.EDG
SAMP4.EDG
SAMP5.EDG
Principal Component Analysis.The eigenvalues of the m components are displayed. Some will be smaller than others; these are more likely to be able to be eliminated than the larger ones. However, it is difficult to predict which ones can be eliminated using this table alone. The next plot, of the individual eigenvectors, gives more information:
Program PCA Version 0.1
Min, max eV to crop data [12599.2, 12748.5] : 12640 12710 (clip the energy range if desired)
Data files :
File 1 SAMP1.EDG
File 2 SAMP2.EDG
File 3 SAMP3.EDG
File 4 SAMP4.EDG
File 5 SAMP5.EDG
------------------------
Component Eigenvalue
1 35.9097
2 3.43393
3 2.86981
4 0.880477
5 1.32538
------------------------
Press 1 to plot Eigenvectors :1The m eigenvectors (principal components), scaled by their eigenvalues, will be displayed (U.V). Note that these principal components are not component spectra, but mathematical constructs. Some will show distinct features, while others may show essentially noise. These latter ones are the candidates for elimination. Based on the eigenvalues and the plots, use the next part of the program to test the elimination of eigenvalues.
Enter GPLOT device number for plot [10] :
Enter route for plot [TT:] :
Press 1 to plot Eigenvectors :One eigenvalue is chosen to be eliminated - in this example #4 was chosen as it had the smallest eigenvalue and its plot looked like noise. After its elimination, the original spectra are reconstructed and the residual sum of squares is calculated for each spectra (see table above). If the eliminated eigenvector is not significant, then all the residuals will be low. However, the absolute value of the residuals will depend on the level of noise in the original spectra. The results can also be examined graphically:
Press 1 to test Eigenvalues, 2 to quit :1
Enter Eigenvalue to eliminate [0] :4
------------------------
4 Eigenvalues
Total Residual : 0.3216760E-02
Component Residual
1 0.2194201E-02
2 0.2202843E-04
3 0.6603940E-03
4 0.2082951E-03
5 0.1318413E-03
------------------------
Press 1 to plot :1Component (original spectrum) 1 was chosen for this plot because it showed the largest residual, and therefore is most likely to show a real (i.e. greater than just the noise) deviation. All n components can be examined in this way. In this case of eigenvalue 4, the deviations are well within the level of the noise, and so this eigenvector is not contributing significantly. The elimination of the next component can then be tested; in our case #5 also is not significant, as judged by the reconstructed spectra. On testing the elimination of #3, however, the following is obtained:
Enter Component to plot [1] :1
Enter GPLOT device number for plot [10] :
Enter route for plot [TT:] :
Press 1 to plot Eigenvectors :and the reconstructed spectra show significant deviations. Thus the elimination should be reversed:
Press 1 to test Eigenvalues, 2 to quit :1
Enter Eigenvalue to eliminate [5] :3
------------------------
2 Eigenvalues
Total Residual : 0.4467924E-01
Component Residual
1 0.3871157E-02
2 0.2383120E-01
3 0.6740484E-03
4 0.4575982E-02
5 0.1172685E-01
------------------------
Press 1 to plot :and, having achieved a reasonable set of components, the program is exited:
Press 1 to undo elimination :1
Data files :
File 1 SAMP1.EDG
File 2 SAMP2.EDG
File 3 SAMP3.EDG
File 4 SAMP4.EDG
File 5 SAMP5.EDG
------------------------
Component Eigenvalue
1 35.9097
2 3.43393
3 2.86981
4 0.000000E+00
5 0.000000E+00
------------------------
Press 1 to plot Eigenvectors :
Press 1 to test Eigenvalues, 2 to quit :2
Thus, from PCA it can be concluded that three components are needed
to fit this set of spectra. However, it gives no information about the
spectral identity of those components. Note also that the elimination of
eigenvalues is somewhat subjective. In the ideal case, with a perfect series
of unknown spectra and no noise, the eigenvalues are rigorously zero. With
the introduction of noise and other experimental uncertainties, these values
are non-zero and whether or not to eliminate them becomes a more subjective
question.
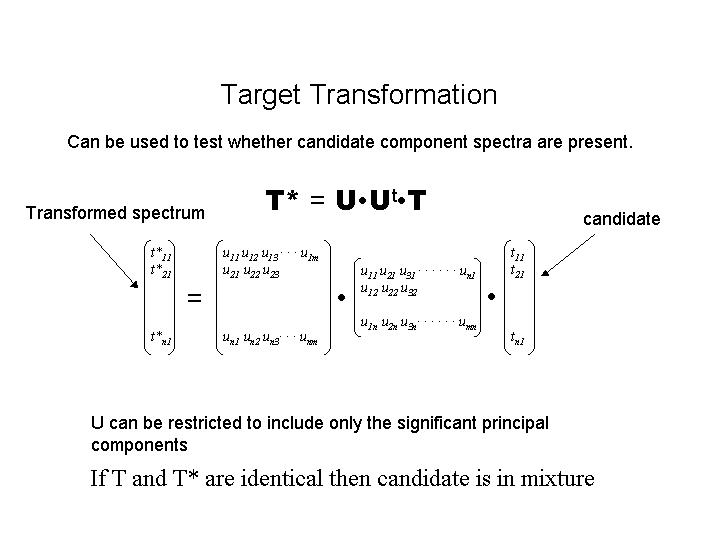
target model1.edgand the model spectrum (T) will be graphically compared with its target transformation (T*). The graphical correspondence of the spectra, together with the residual value (displayed on the plot) can be used to test whether the spectrum is a component. Note that again this is a subjective test.
Please refer to the documentation on Edge
Analysis for more information on DATFIT.